How and why does multidrug resistance evolve?
Rosie Clover is studying for a PhD as part of the BBSRC Doctoral Training Partnership
A love of all things microbial is what drove me to the University of Manchester, where I began my studies as a MSc Medical Microbiology student. We were taught how to identify and test bacteria in a number of different ways that would allow us to identify causes of infection and find the best route to treating it. During my Dissertation Project I was given the opportunity to work with Dr Danna Gifford and Dr Chris Knight to phenotypically characterise and develop antibiotic resistance profiles for 100 clinical multidrug resistant strains of E. coli collected from Greater Manchester. It was fascinating to see how the same species of bacteria can grow and (mis)behave so differently and I found myself wanting to know as much as I possibly could about this collection of strains.
My MSc dissertation set the foundations for my PhD project which is focused on antibiotic resistance evolution in this clinical collection of E. coli. Coming into the Knight/Gifford lab, I knew very little about evolution and its application to antibiotic resistance. I realised very fast that antibiotic resistance extends further than I had previously thought and we need to know more about the how and why of resistance, particularly multidrug resistance in clinical settings. At the start of my project, I began by looking at how acquiring multiple resistance genes or mutations impacts the strain’s ability to grow in an antibiotic-free environment. The acquisition of resistance is usually associated with an impaired ability to grow; a concept known as the ‘cost of resistance’. As I read up on the cost of resistance, I found that most of the current literature focused on single-drug resistance or laboratory strains. So, how does the cost of resistance change for clinical strains that are already multidrug resistant?
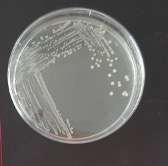
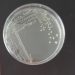
To find out whether the cost of resistance exists among a multidrug resistance clinical collection of E. coli, we looked at resistance on a phenotypic and genotypic level. We began by exposing them to a series of clinically relevant antibiotics. They were grown on antibiotic-containing agar to provide a definite yes/no to growth in presence of antibiotics. They were then sent for whole genome sequencing (WGS) and their genomes analysed to develop a profile of resistance genes and mutations carried by each strain. (Which for a girl who had only ever used her laptop for Microsoft Word, was a huge learning curve!) By doing this, we could compare the growth of the E.coli in an antibiotic-free environment to the number of resistances it carried both phenotypically and genotypically. Between these two methods of characterising resistance, we observed no negative impact of carrying resistances on the strain’s growth in an antibiotic-free environment. Thinking about what this means, it implies that terminating antibiotic therapies may not produce the intended decline in prevalence of strains resistant to those antibiotics. As such my data poses more question that answers! Questions which as an AMR community we work together to answer.
What is next for my PhD? I know which antibiotics these strains are resistant to and how they grow in different environments: Using these well-characterised strains I can next begin testing how resistance evolves in each of these strains as a function of their pre-existing resistance and their fitness. The next stage of my project therefore will expose a subset of the strains to slowly increasing concentrations of antibiotics to quantify the rate and mode by which these clinical strains gain new resistances. By moving my research in this direction we will be able to see how resistances are maintained and multidrug resistance develops in clinical populations of E.coli.

0 Comments